Genome contact map browser¶
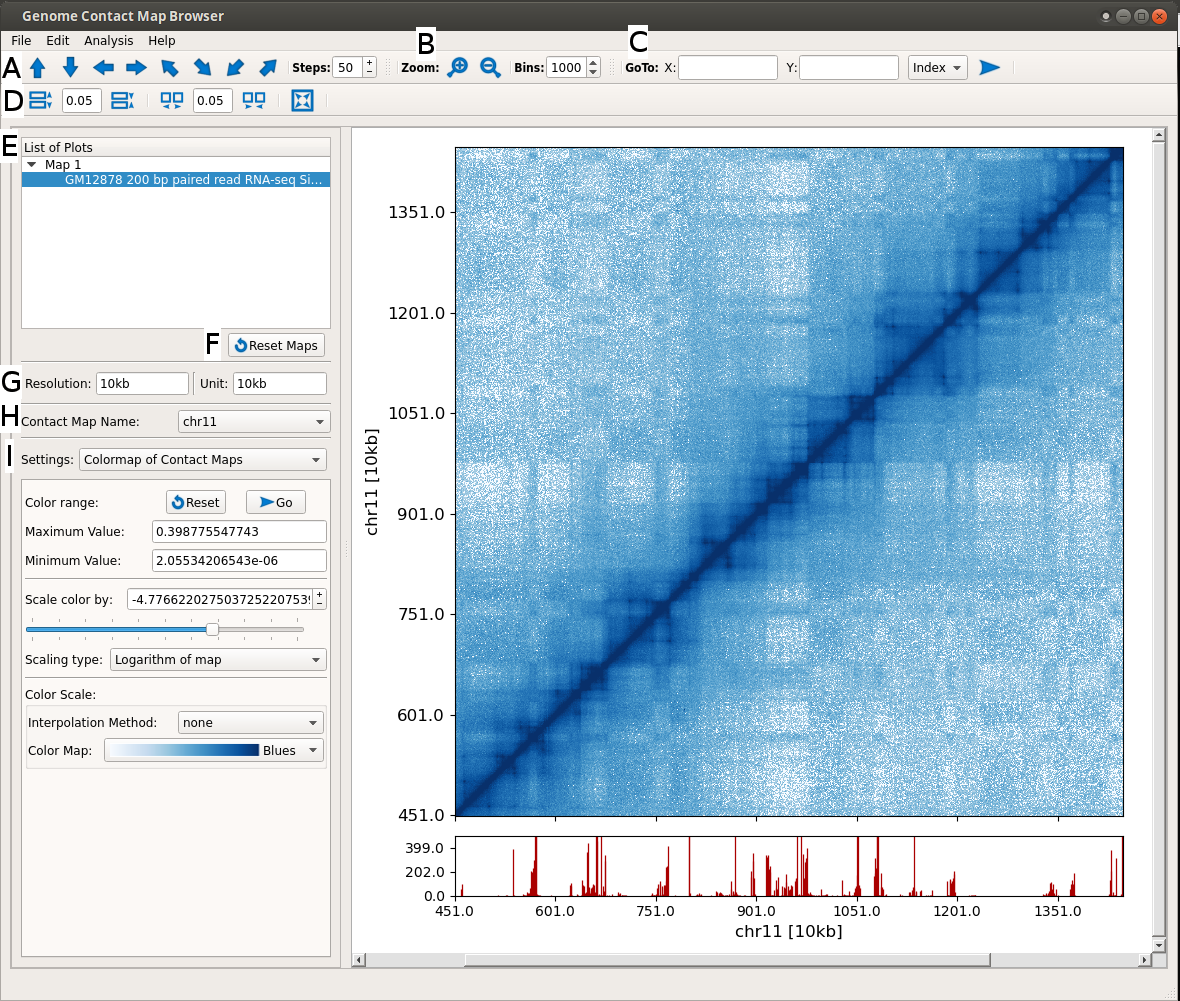
It is an application to browse genome contact maps along with respective genomic track datasets. It acts as an interactive visualizer, where user can browse the data by dragging and zooming.
- To launch browser, execute following command:
gcMapExplorer browser
- Click on arrow to move the maps in respective directions. The genomic datasets also follow automatically.
- Click on these buttons to zoom in and out. The genomic datasets also follow automatically.
- Use this to go to on specific coordinate. Input can be real or indexed coordinate.
- Change width-spacing between plots.
- List of all plots. Active plot is selected. One can select a plot to make it active.
- Reset all maps to original view.
- Display information of current resolution and unit shown in plots.
- Name of contact map. Usually chromosome name.
- Various options. Presently shown for color mapping and scaling of maps. It can be used to modify colormaps, color scaling range and color scaling type.
Add a genomic track dataset¶
Click on File -> Add genomic dataset to ... and select the contact map
for which new dataset is need to be added. A new window will appear where user
may select a compatible file by clicking on Open button. Afterwards,
different box will appear depending on the input file format.
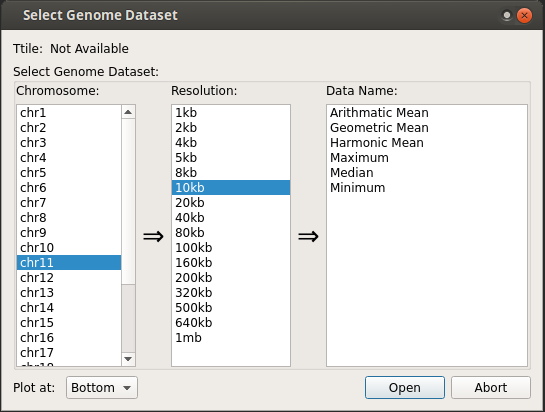
h5 file: User is prompted to select the dataset.
By default, the genomic track is added at the bottom of map. However, one can select
topin the above dialog box to plot the track at the top of the contact map.bigWig/wig/bed file : Since, only h5 files are compatible with browser, a window will appear as similar to h5Converter to convert these files to h5 on the fly. By default, the generated h5 file will be deleted after use because
Removeis checked. To save the generated h5 file, uncheckRemoveoption and change location of output file.Once dataset is converted, user is prompted to select the dataset as shown above for h5 input file.
To reduce the waiting time for conversion process, only dataset for the required chromosome is converted. When later another chromosome is selected in
browser, h5Converter will again appear and dataset will be again converted for the selected chromosome. Once dataset for a chromosome is converted, it remains stored in the h5 file. Therefore, when this chromosome is re-selected inbrowser, no conversion is required and dataset is plotted inbrowserinstantly.Note
The options in h5Converter are only editable at first time. Later when another chromosome is selected in
browser, this box will again appear, however, options will not be editable.
Change page size and orientation¶
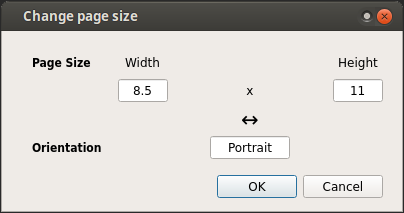
Click on Edit -> Change Page Size and select desired page size. In case of
custom page size, click on Custom. A dialog will open, where user can specify
new page size.
Save as image¶
Click on File -> Save Plot, choose file name with acceptable
extension and click on Save button to save the plot as image file.
Change Genomic track plot setting¶
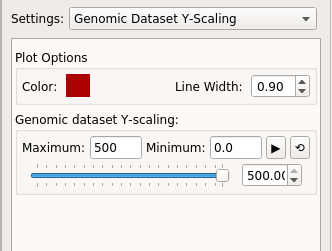
Select a Genomic Dataset Y-Scaling option in Settings at right panel.
Below several options will appear. These include color, line width, maximum and
minimum limit along Y-axis.
Change marker setting¶
Select a Marker option in Settings at right panel. Below options will
appear to modify the marker settings.
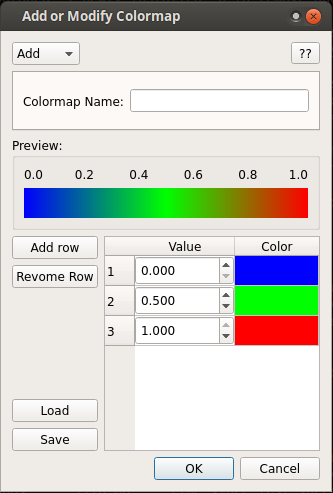
User defined colormap¶
Although several colormap is already included in the browser. One may generate
own colormap and later modify it using the implemented option. To open this, click
on Edit -> Add/Modify colormap.
This box can be used to create or modify the colormaps. The shown colormap can be saved as a text file for later use.
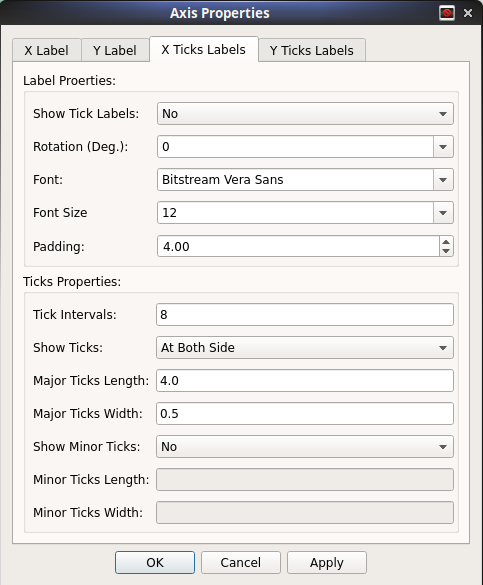
Modify Axis Properties¶
Properties of both X and Y axis are highly customizable. User may customize most of the properties such as font, tick-lengths, axis-labels etc. To open this box, right click on plot and choose the axis properties.
Save and Load View Points¶
Visualization state can be also saved as view-points. To add a view Points,
select View Points in settings at left panel. Then, click on Add
buttons. It will add the current visualization state in the list. One can add
as many as view points in the list.
Each view-point store all settings of the plot. By double-clicking on a view-point, it can be loaded as it was stored.
More importantly, all these view-points can be saved in a file for later viewing. Click on
File -> Save Visual State and View Points to save exact visualization states and
all view-points as a file.
The saved file can be opened by clicking on File -> Load Visual State and View Points. The file can be
also loaded directly by following command.
gcMapExplorer browser visual_states.gvs
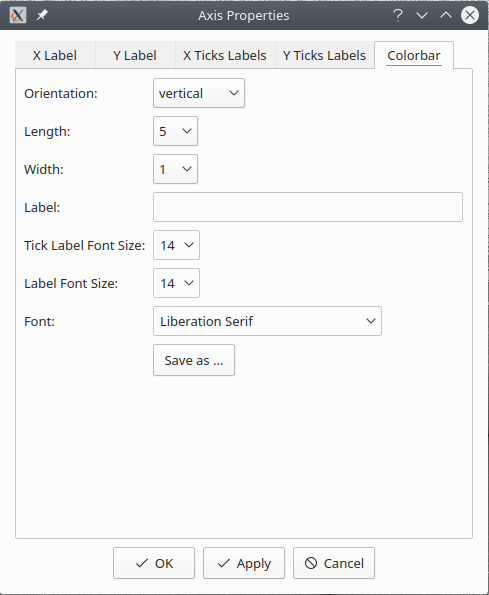
Save color-bar¶
The browser does not display a color-bar. However, a color-bar can be saved as an image
file, separately. To save the color-bar, right click on map and select Save colorbar....
Input the information, and save the color-bar as image file.