Using masked array with Hi-C map¶
Large Hi-C map data contains missing values. During analysis, sometimes we need to ignore these values. numpy.ma module can be used to perform mathematical operations after masking these missing values.
See also
Module numpy.ma
At first, we import modules:
- gcMapExplorer.ccmap for loading
.ccmapfiles - numpy for array operations
- numpy.ma for masked array module
- scipy.stats.mstats for correlation calculation using masked array
In [1]:
import gcMapExplorer.ccmap as cmp
import numpy as np
import os
from numpy import ma
from scipy import stats
import matplotlib.pyplot as plt
# To show inline plots
%matplotlib inline
plt.style.use('ggplot') # Theme for plotting
To calculate correlation between Hi-C maps¶
Load two Hi-C data
In [2]:
ccmapOne = cmp.load_ccmap('output/CooMatrix/normalized/chr15_100kb_normKR.ccmap', workDir=os.getcwd() )
ccmapOne.make_readable()
ccmapTwo = cmp.load_ccmap('output/CooMatrix/normalized/chr20_100kb_normKR.ccmap', workDir=os.getcwd())
ccmapTwo.make_readable()
Determine smallest shape
Matrix size can be different, therefore smallest size is determined to calculate element-wise correlation.
In [3]:
if ccmapOne.shape[0] <= ccmapTwo.shape[0]:
smallest_shape = ccmapOne.shape[0]
else:
smallest_shape = ccmapTwo.shape[0]
Generate masks
- At first, generate mask from two matrices sepratealy, and then combine it.
- Also, use slicing operations to derive subset of matrix with smallest shape.
- During masking, lower-triangular part is also masked with diagonal offset of five. Because the values at Hi-C map diagnoal is usually large, correlation could be high due to these large values. Moreover, we are usually intereseted in off-diagonal regions of the Hi-C maps.
In [4]:
m1 = ccmapOne.matrix[:smallest_shape, :smallest_shape] <= ccmapOne.minvalue # Mask all minimum values
m2 = ccmapTwo.matrix[:smallest_shape, :smallest_shape] <= ccmapTwo.minvalue # Mask all minimum values
mask = ( m1 | m2 ) # Combine both masks
mask[np.tril_indices_from(mask, k=5)] = True # Also, Mask lower-triangle of matrix with five diagonal offset
Warning
For huge matrices, above operations could be memory/RAM consuming and script may crash.
See also
Generate masked matrix
Now, using numpy.ma module, generate masked matrices
In [5]:
maskedMatrixOne = ma.array(ccmapOne.matrix[:smallest_shape, :smallest_shape], mask=mask)
maksedMatrixTwo = ma.array(ccmapTwo.matrix[:smallest_shape, :smallest_shape], mask=mask)
Calculate correlation coefficiant
- Pearson correlation coefficiant using scipy.stats.mstats.pearsonr
In [6]:
corr, pvalue = stats.pearsonr(maskedMatrixOne.compressed(), maksedMatrixTwo.compressed())
print('Pearson Correlation: ', corr)
corr, pvalue = stats.spearmanr(maskedMatrixOne.compressed(), maksedMatrixTwo.compressed())
print('Spearman Correlation: ', corr)
Pearson Correlation: 0.50507
Spearman Correlation: 0.358783199081
Crorrelation using gcMapExplorer.cmstats module¶
Shown above is a simple step-by-step example to calculate
correlation-coefficiant using masked array. However, above method may
fail in case of huge matrices. Therefore, use the implemented function
gcMapExplorer.cmstats.correlateCMaps function
See also
- About
gcMapExplorer.cmstats.correlateCMaps()in more details
In [7]:
from gcMapExplorer import cmstats
corr, pvalue = cmstats.correlateCMaps(ccmapOne, ccmapTwo, diagonal_offset=5)
print('Pearson Correlation: ', corr)
corr, pvalue = cmstats.correlateCMaps(ccmapOne, ccmapTwo, corrType='spearman', diagonal_offset=5)
print('Spearman Correlation: ', corr)
Pearson Correlation: 0.50507
Spearman Correlation: 0.358783199081
Both above shown step-by-step example and implemented functions yielded similar correlation values.
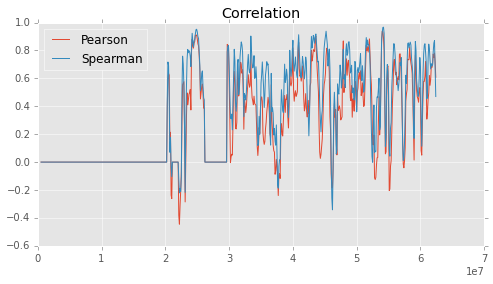
Block-wise Crorrelation¶
To identify local difference between two maps, block-wise coorelation could be more helpful. A block is created and slided along the diagonal, and for each new position, correlation is calculated.
In [8]:
# Pearson correlation
pearson, p_centers = cmstats.correlateCMaps(ccmapOne, ccmapTwo, diagonal_offset=2, blockSize='1mb',
slideStepSize=1, outFile='pearson.txt', workDir=os.getcwd())
# Spearman correlation
spearman, s_centers = cmstats.correlateCMaps(ccmapOne, ccmapTwo, corrType='spearman', diagonal_offset=2,
blockSize='1mb', slideStepSize=1, outFile='spearman.txt',
workDir=os.getcwd())
INFO:correlateCMaps: Block-wise correlation with [1mb] block-size
INFO:correlateCMaps: Number of Blocks: 63
INFO:correlateCMaps: Size of each Block in bins: 10
INFO:correlateCMaps: Number of Overlapping bins between sliding blocks: 9
INFO:correlateCMaps: Block-wise correlation with [1mb] block-size
INFO:correlateCMaps: Number of Blocks: 63
INFO:correlateCMaps: Size of each Block in bins: 10
INFO:correlateCMaps: Number of Overlapping bins between sliding blocks: 9
In [9]:
# Plot the values for visual representations
fig = plt.figure(figsize=(8,4)) # Figure size
ax = fig.add_subplot(1,1,1) # Axes first plot
ax.set_title('Correlation') # Title first plot
ax.plot(p_centers, pearson, label='Pearson') # Plot for Pearson correlation
ax.plot(s_centers, spearman, label='Spearman') # Plot for Spearman correlation
ax.get_xaxis().get_major_formatter().set_useOffset(False) # Prevent ticks auto-formatting
plt.legend(loc=2)
plt.show()
In [10]:
path='/home/rajendra/workspace/genome_3d_organization/GM12878_CellLine/HiCmapDataPyObject/resolution_10kb/'
ccmapsOne, ccmapsTwo = [], []
for i in range(1, 23):
ccmapsOne.append(path+'Chr{0}_NormKR.hicmap'.format(i))
for i in range(1, 23):
ccmapsTwo.append(path+'Chr{0}_Rawobserved.hicmap'.format(i))
for i in range(len(ccmapsOne)):
ccmapOne = cmp.load_ccmap(ccmapsOne[i], workDir=os.getcwd())
ccmapOne.make_readable()
ccmapTwo = cmp.load_ccmap(ccmapsTwo[i], workDir=os.getcwd())
ccmapTwo.make_readable()
print(cmstats.correlateCMaps(ccmapOne, ccmapTwo, corrType='spearman', diagonal_offset=5))
del ccmapOne
del ccmapTwo
(0.74487690967318498, 0.0)
(0.74063722421215261, 0.0)
(0.75637101395441264, 0.0)
(0.75799064378987224, 0.0)
(0.76385777207989702, 0.0)
(0.7581134503870115, 0.0)
(0.77085117905282485, 0.0)
(0.78712169997619941, 0.0)
(0.75681100168481319, 0.0)
(0.79607602070672256, 0.0)
(0.79340494862540256, 0.0)
(0.79611875779812202, 0.0)
(0.78246725829489128, 0.0)
(0.81078979072455271, 0.0)
(0.82796003802636897, 0.0)
(0.81646908774859528, 0.0)
(0.8513992946693909, 0.0)
(0.83321467022045526, 0.0)
(0.89497615144311282, 0.0)
(0.89713225129243968, 0.0)
(0.86860067840860633, 0.0)
(0.87945995663009324, 0.0)