Genome Contact Map Explorer - gcMapExplorer¶
It is a platform to visualize and analyze the contact maps that are generated from Hi-C experiments. This package is developed by considering the huge size of contact maps at very fine resolution. It contains
- Graphical User Interface Applications - Several windows like applications to perform tasks.
- Command Line Interface - Several commands to perform tasks.
- Application Programming Interface - It can be used to perform analysis by any mathematical operations through programming.
For Discussion and Questions, visit this forum
Features¶
Support for huge contact maps - Use of Disk instead of RAM - Matrices/arrays are stored in Disks - mathematical operations by directly reading/writing from/to Disks, without loading them into RAM
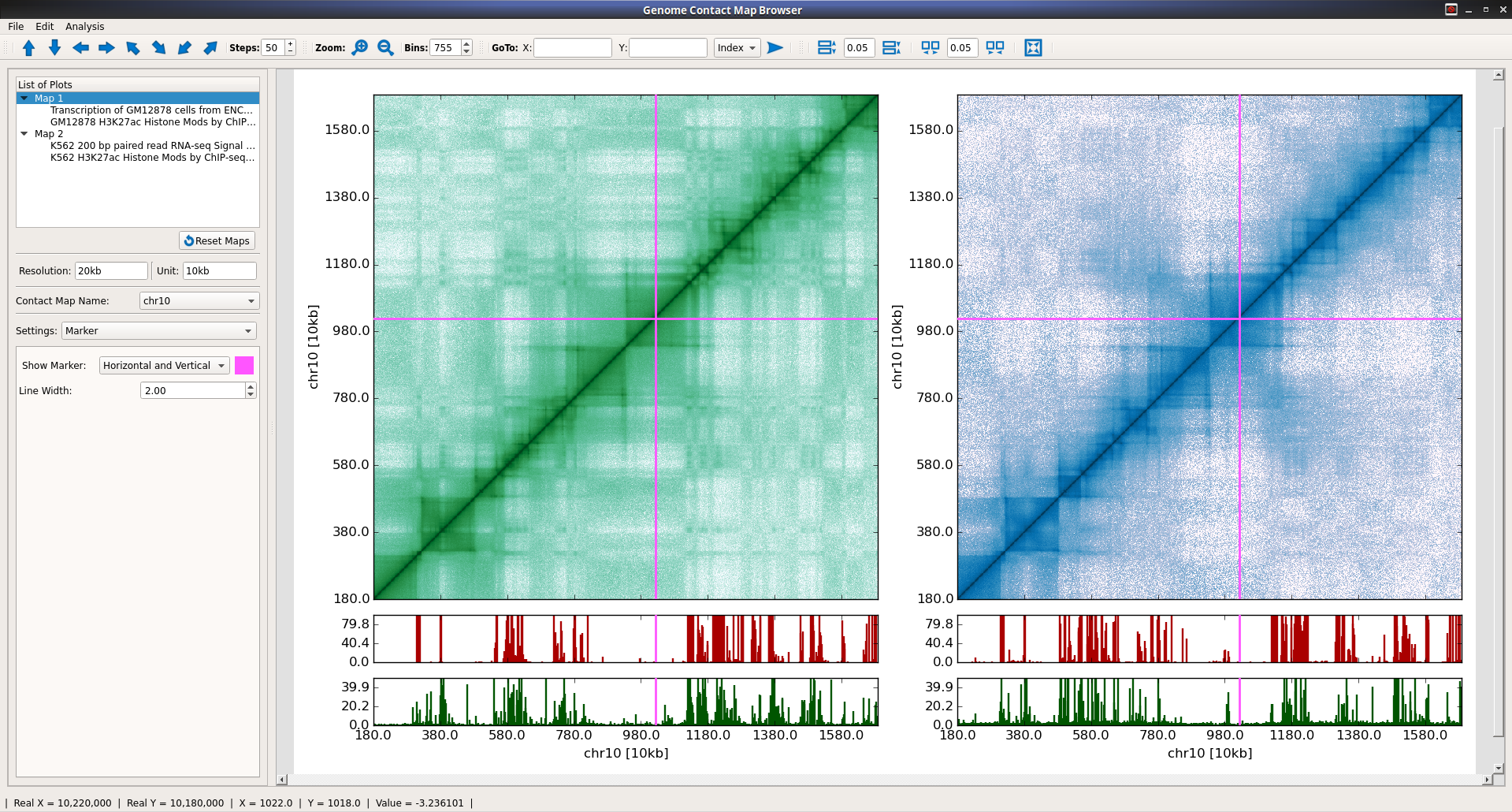
A browser with rich interfaces for Comparative and Interactive visualization of two dimensional contact maps along with genomic datasets such as produced by DNase-seq, ChIP-seq, RNA-seq etc.
Contact maps can be zoomed in/out from finest resolution to whole chromosome level.
Rich customizations of color scale for contact maps visualization
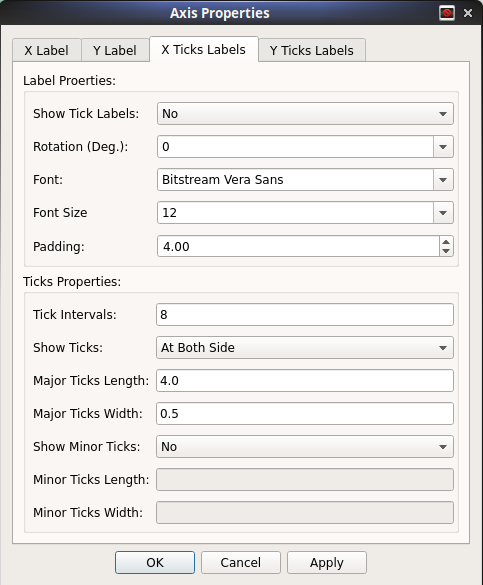
Rich customizations of X- and Y- axis properties.
- Normalization of contact maps by
- Iterative Correction (IC)
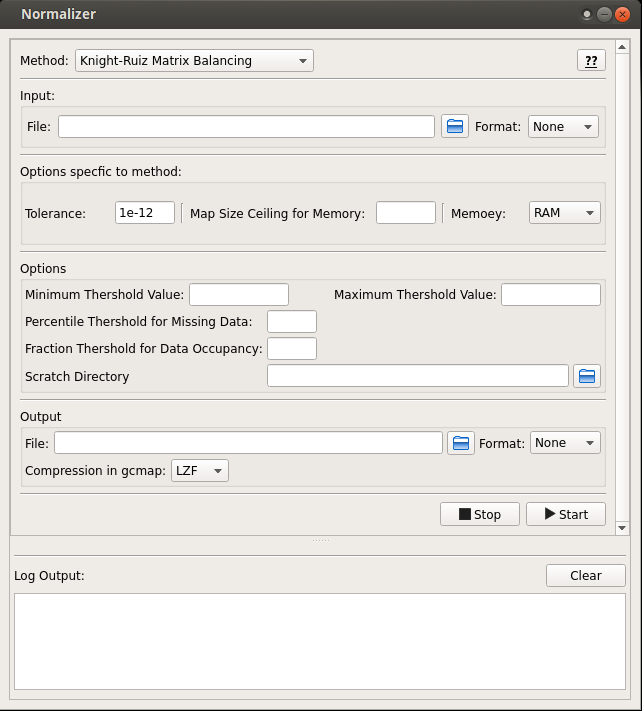
- Knight-Ruiz Matrix Balancing (KR)
- Vanilla-Coverage (VC)
- Distance-Frequency
A new file format based on HDF5 for genome contact map and genomic track datasets.
- Portable, platform independent and can be read through C/C++, JAVA, Python and R programming language.
- Very fast to read - fast browsing of contact maps and genomic datasets
Another file format for chromosomal contact map - much faster than above format to read/write but not compact. Suitable for performing calculations.
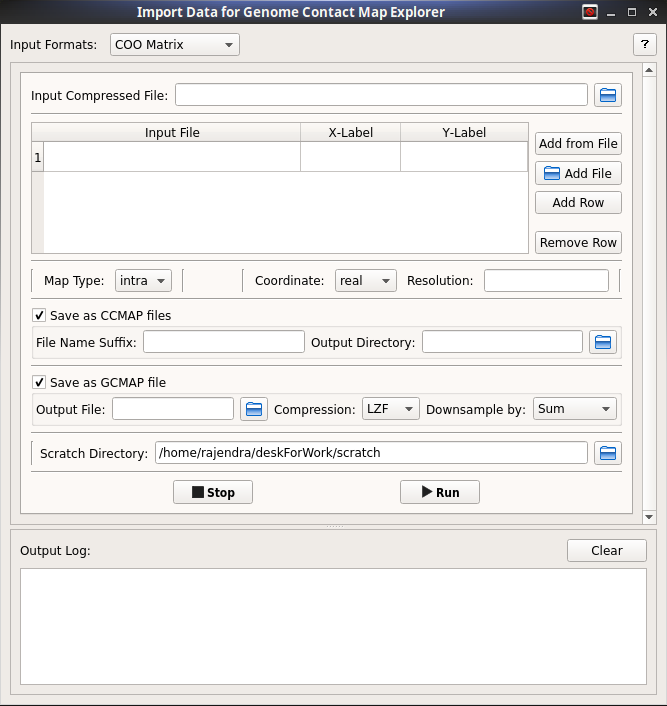
A GUI interface and commands to convert Coordinate Sparse, Pair Coordinate Sparse, HOMER Interaction matrix, Bin-Contact formats into the new gcmap and ccmap formats.
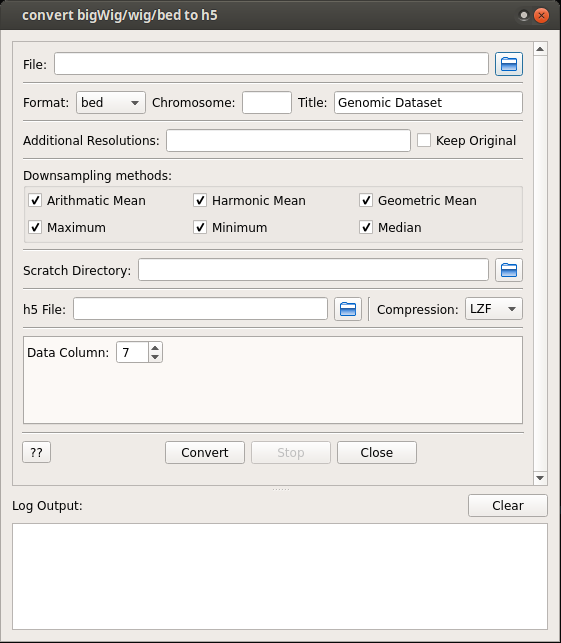
Interface and commands to convert bigWig/wig/bed file to genomic track dataset h5 file.
Publication ready images at one click.
Citation¶
Rajendra Kumar, Haitham Sobhy, Per Stenberg and Ludvig Lizana. Genome Contact Map Explorer - A platform for the comparison, interactive visualization and analysis of genome contact maps. Nucleic Acids Res. (2017).
Screen-shots¶
Contents¶
- Requirements and Installation
- How to use gcMapExplorer?
- Genome Contact Map Browser
- About file formats
- Normalization of Hi-C maps
- Frequently asked questions
- Download example datasets
- Summary of Python Modules
- Examples using Python Modules
- How to Import external Hi-C map data?
- How to normalize Hi-C map?
- How to access Hi-C data from ccmap file?
- Using masked array with Hi-C map
- How to export Hi-C data as text file?
- How to access map from gcmap file?
- Comparison - original KR algorithm and implemented method
- Comparison - original ICE algorithm and implemented method
- Python Modules documentation